xGen™ Hybridization Capture Core Reagents NEW
Research reagents for the xGen NGS Hybridization Capture workflow
xGen Hybridization Capture Core Reagents provide reliable targeted sequencing functionality across a range of panel sizes and multiplexing levels. They work with xGen Hyb Panels for a complete, high quality target enrichment solution.
xGen NGS—made to maximize.
Ordering
- Generate targeted NGS data with a quick and easy workflow
- Automation-friendly protocol containing no heated buffers, 1X buffers and 1 hour hybridization enables various levels of throughput
- Achieve uniform coverage and reliable capture functionality no matter your desired capture time or input across a range of xGen Hyb Panels
- xGen Universal Blockers effectively block a variety of index adapter designs with proprietary sequence modifications improving on-target results
- xGen 2X HiFi PCR Mix now included, so no need to source components from multiple vendors
Transform Your NGS Workflow with Automation
Looking to streamline your NGS workflows? Discover how automation can enhance efficiency and consistency in your lab with our NGS Automation solutions.
For customers using the legacy xGen Lockdown reagents, these products can be ordered here.
For customers using the legacy xGen Hybridization and Wash Kit, these products can be ordered here.
Product details
The xGen Hybridization and Wash v3 Kit is designed for use with xGen Hyb Panels and xGen Universal Blockers. This kit consists of two core components—the xGen Hybridization & Wash v3 Reagents and the xGen Hybridization & Wash v3 Beads—to perform the hybridization capture workflow. The latest version of the kit contains a new workflow and components that require no heated buffers or extra dilution steps of buffers to provide a complete, high-quality hybridization capture solution for targeted NGS research that you can complete in one day. The new buffer system allows for hybridization times as short as 1 hour and inputs as low as 100 ng.
Prevent adapter cross-hybridization by using xGen Universal Blockers in your hybridization capture reactions. Adapter sequences are ligated to all library fragments, both on-target and off-target, before enrichment. These adapter sequences can hybridize with each other during enrichment, creating a "daisy-chain" effect, with off-target fragments being captured alongside on-target fragments, which can impact sequencing efficiency.
xGen Universal Blockers bind to platform adapter sequences on a designated strand (usually the inverse of the synthetic adapter) to help prevent non-specific binding. Blocking the adapter sequences significantly improves capture results [1, 2].
Product data
Reduce hybridization capture time and maintain data quality
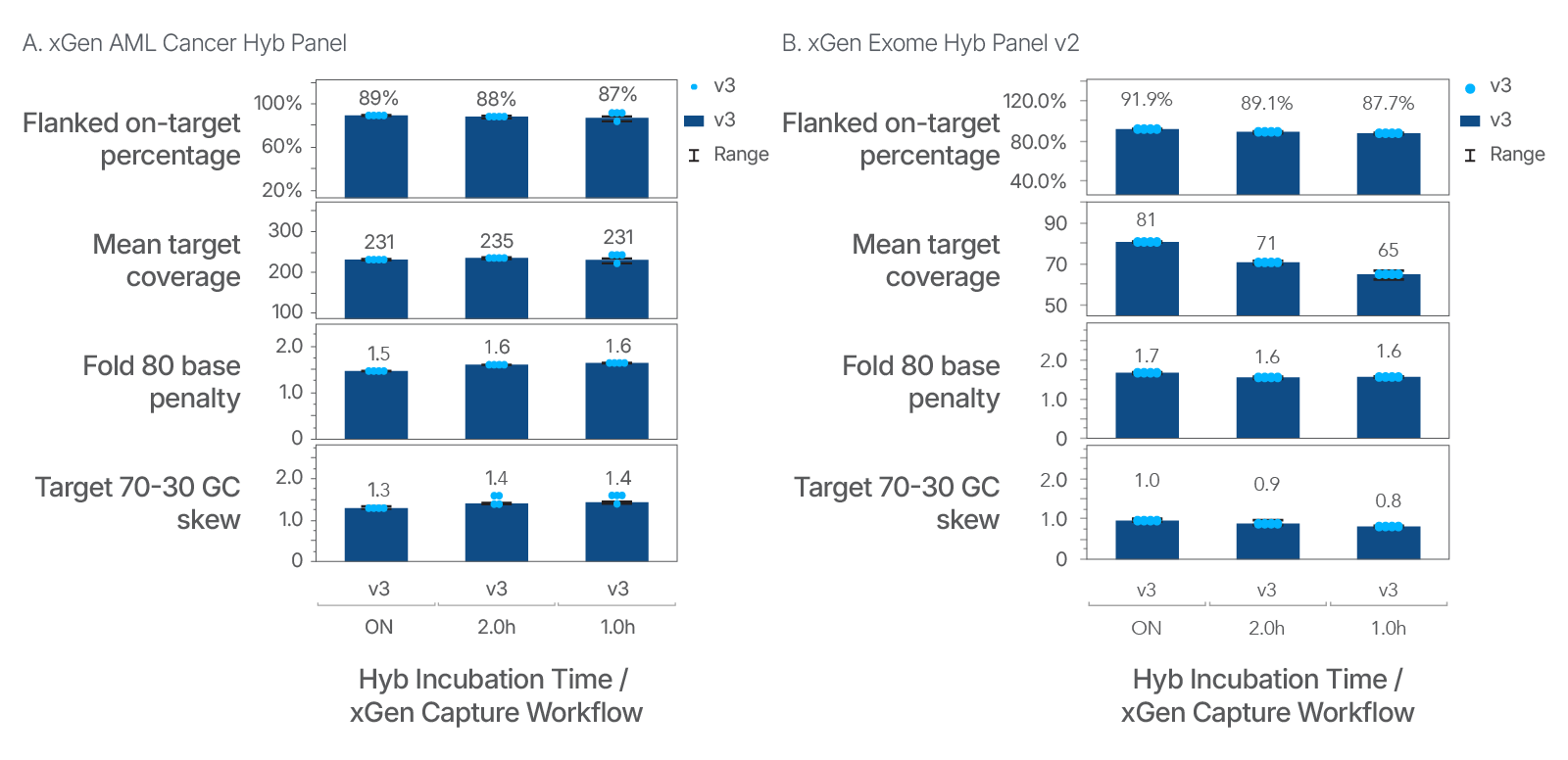
Figure 1. Reduce hybridization capture time and maintain data quality. To evaluate the performance of the new buffer system in xGen Hybridization and Wash Buffer v3, we set up a series of experiments to assess hybridization time with two different panels. The DNA libraries were prepared using xGen DNA EZ Library Prep kit with 25 ng Coriell gDNA NA12878. The DNA libraries were subsequently enriched using 400 ng of input with either the (A) xGen AML Cancer Hyb Panel or (B) xGen Exome Hyb Panel v2 for either overnight (ON), 2 hours, or 1 hour.
Captured libraries were sequenced on a NextSeq 2000 system (Illumina®) and subsampled to 5 M total reads per sample. A standard analysis pipeline was run to produce Picard metrics. The flanked-on target percentage and fold 80 base penalty was consistent for both panels between hybridization incubation times. (A) The mean target coverage and the GC skew were consistent for the xGen AML Cancer Panel. (B) A variation in the GC skew metric was observed between the different hybrization incubation times using the xGen Exome Hyb Panel v2.
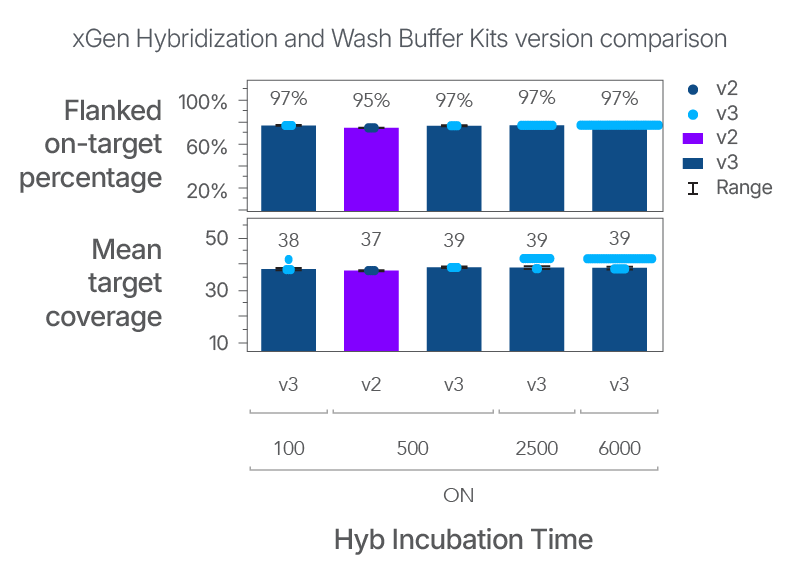
Robust capture across a range of library inputs
Figure 2. Consistent capture between xGen Hybridization and Wash Buffer v2 and v3 across a range of library inputs. To evaluate the performance of the v3 buffer system in xGen Hybridization and Wash Buffer v3 was compared to the xGen Hybridization and Wash Buffer v2 with different library inputs. The DNA libraries were prepared using the xGen cfDNA & FFPE DNA Library Prep v2 MC kit with 25 ng Coriell gDNA NA12878. The DNA libraries were subsequently enriched using 100 ng, 500 ng, 2500 ng, or 6000 ng of library input with the xGen Exome Hyb Panel v2 for overnight hybridization. Captured libraries were sequenced on a NextSeq 2000 system (Illumina®) and subsampled to 30 M total reads per sample. Standard analysis pipeline was run to produce Picard metrics. Despite the low sequencing data (30 M reads), the mean target coverage was consistent and robust across conditions.
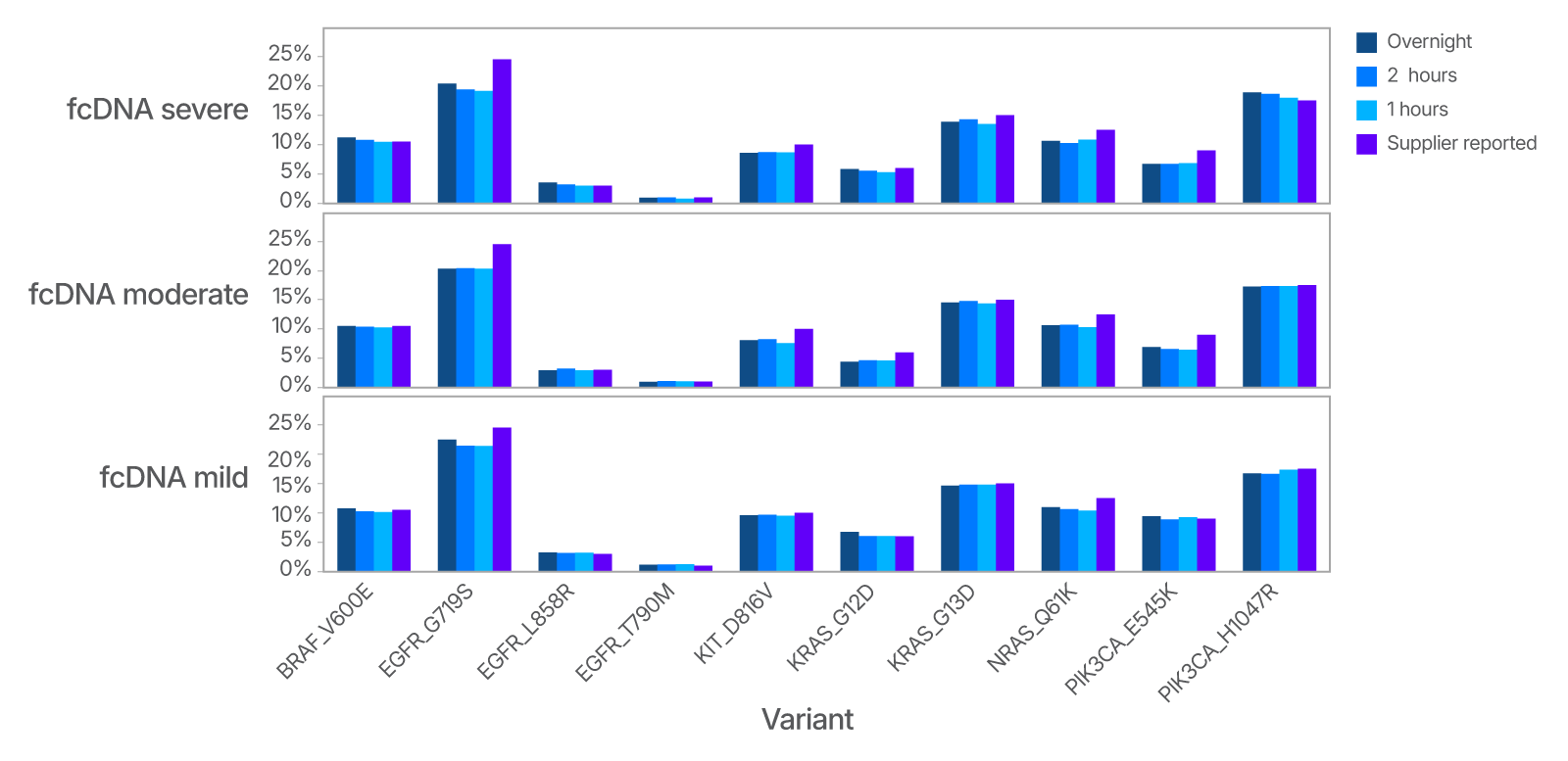
Detect SNVs even in degraded sample types
Figure 3. Impact of hybridization time on SNV detection in degraded samples. Three commercially available samples with known SNVs at measured frequencies, and with differing degrees of DNA damage (as measured by DIN score), were used as test library prep input. One hundred nanograms of DNA from each sample was used with the xGen cfDNA & FFPE DNA Library Prep Kit following IDT’s published protocol. Each sample was prepared in quadruplicate. Five hundred nanograms of library was used in a singleplex capture for various times of hybridization (overnight, 2 hours, and 1 hour). Samples were sequenced to approximately 100 M total reads and analyzed, to create ssConsensus data that was interpreted by VARDICT. The measured allele frequency for each of the known SNVs is indicated for different hybridization times. The following samples were used: Horizon discovery standards - HD803 (fcDNA severe), HD799 (fcDNA moderate), and HD798 (fcDNA mild).
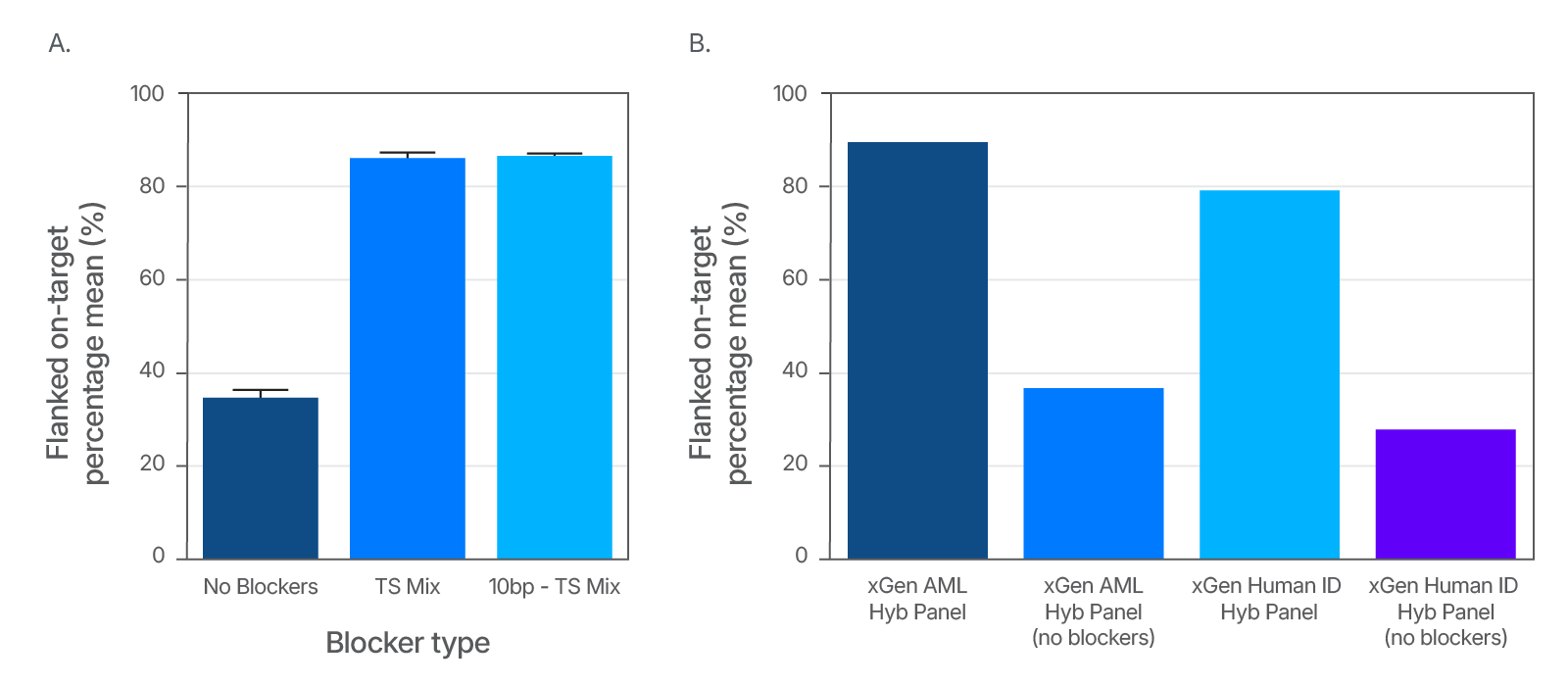
Increase on-target rates by using xGen Universal Blockers
Figure 4. An example of improved on-target rate delivered by xGen Universal Blockers across Illumina TruSeq® (A) and Illumina Nextera® (B) adapter styles. (A) 1 µg input DNA libraries were prepared from human genomic cell line NA12878 (Coriell) using either 8 nt or 10 nt adapters and enriched using the xGen AML Cancer Hyb Panel (single-plex, n = 3 for each condition) with xGen™ Universal Blockers TS, xGen™ Universal Blockers 10bp TS, or without blockers. Sequencing was performed on a NextSeq 500 System (Illumina) to generate 2x150 bp paired-end reads. (B) 50 ng cell line NA12878 (Coriell) gDNA was used for Nextera library preparation. Amplified libraries were enriched in 4-plex reactions (n = 3) using the xGen AML Cancer Hyb Panel, or in 12-plex (n = 1) using the xGen Human ID Hyb Panel, with xGen Universal Blockers NXT or without blockers (n = 1). Sequencing was performed on a NextSeq System (Illumina) to generate 2x75 bp paired-end reads. Libraries and captures from both figures were amplified using KAPA 2X HiFi PCR Mix, as this data was generated prior to release of xGen 2X HiFi PCR Mix.
High on-target and uniform coverage with xGen Universal Blockers 10bp TS
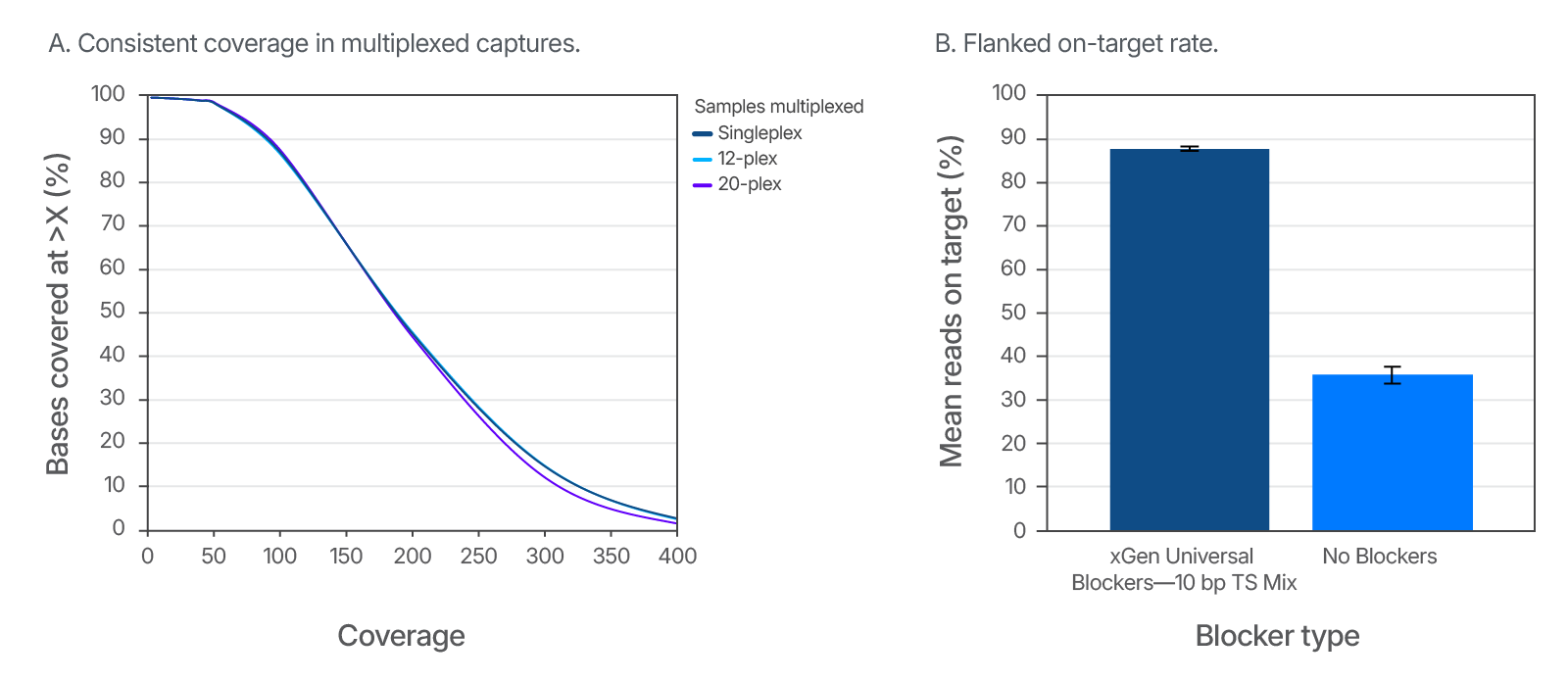
Figure 5. Consistent coverage in multiplexed captures and improved on-target rates with the addition of xGen Universal Blockers. (A) Example of consistent results with multiplexed samples using the xGen Universal Blockers 10bp TS. DNA libraries (500 ng input per sample) from cell line NA12878 (Coriell) were enriched in single (n = 8) or multiplex reactions (n = 3) using the xGen AML Cancer Hyb Panel. The libraries were created using 10 bp TruSeq adapters with unique dual indexes. Sequencing was performed on a NextSeq 500 System (Illumina) with 2x150 bp paired-end reads. Coverage was consistent across all multiplexing levels tested. (B) Example of improved on-target rates delivered by xGen Universal Blockers 10bp TS. DNA libraries (500 ng input per sample) from cell line NA12878 were enriched in single-plex reactions (n = 3) using the xGen AML Cancer Hyb Panel with and without xGen Universal Blockers 10bp TS. Sequencing was performed on a NextSeq 500 System to generate 2x150 bp paired-end reads. On-target values (with 150 bp flank) were averaged across experiments. Libraries and captures from both figures were amplified using KAPA 2X HiFi PCR Mix, as this data was generated prior to release of xGen 2X HiFi PCR Mix.
Resources
Frequently asked questions
Would a reaction in the xGen™ Hybridization Capture and Wash v3 Kit protocol fail if the sample didn't reach room temperature after each heated step or was held too long at room temperature?
Samples held at room temperature for a slightly longer time beyond the recommended two minutes had no observable impact. Captures have also been tested by moving on immediately without allowing the capture to cool sufficiently without observable impact.
Is the xGen™ Pre-hybridization Normalase™ Module compatible with the xGen Hybridization and Wash v3 Kit?
Yes, xGen Pre-hybridization Normalase serves as an extension of the library preparation stage by normalizing samples for xGen Hybrid Capture.
Can I multiplex libraries into a single hybrid capture when using the xGen™ Hybridization and Wash v3 Kit?
Yes, multiplexing of sample libraries is supported. The sum total recommend library input should not exceed 6 µg. The total input range can be 100 ng to 6 µg. If multiplexing, ensure indexes are not repeated and have an edit distance of at least 3.
Do I need to prepare the buffers in any way before performing captures with the xGen™ Hybridization and Wash v3 Kit?
Allow the reagents to come to room temperature. Buffers do not need to be preheated to a high temperature for use.
What do I do if I get variable results between replicate captures when using the xGen™ Hybridization and Wash v3 Kit?
A likely root cause is inadequate mixing of the hybrid captures. Thorough mixing is vital for consistent results. Mixed samples should be visually inspected to ensure homogeneity.
Can I use my old xGen™ panels that were used with the xGen Hybridization and Wash v2 Kit with the v3 Kit?
Yes, xGen Hybridization and Wash Buffer v3 Kit is designed to work with any standard xGen panels, stocked or custom.
How do I deal with a spike in panel when using the xGen™ Hybridization and Wash v3 Kit?
Option 1: Panels can be mixed together, dried down and resuspend as a new panel.
Option 2: A user can increase the volume from 4 µL to 6 µL (the spike-in volume) of the added xGen panel and then increase the volume of the xGen Hybridization reagent from 14 µL to 15 µL, rendering the total volume to 21 µL of capture reaction.
What if I don’t have a Speed Vac to dry down libraries needed for the xGen™ Hybridization and Wash v3 Kit?
Please refer to Appendix A of the xGen Hybridization and Wash v3 Kit protocol for the AMPure XP Bead DNA concentration method. A user can also consider an oven/incubator at 65°C for a few hours.
Are there safe stopping points with the xGen™ Hybridization and Wash v3 Kit protocol like the previous versions?
Yes, there are safe stopping points, please follow the protocol as written. The safe stopping points allow a user to pause and return to the protocol. Various temperatures are possible depending on the specific step.
When using the xGen™ Hybridization and Wash v3 Kit, can I hybridize with a thermocycler lid temperature below 105°C?
Yes. We recommend setting thermocycler lid at least 10°C above the sample temperature for all heated steps.
When using the xGen™ Hybridization and Wash v3 Kit, can I hybridize at 65°C instead of 70°C?
We strongly recommend hybridization at 70°C, especially for large panels and high library inputs. Experienced users and investigators can further optimize and tailor the conditions for their specific needs.
Can I substitute buffers or reagents from versions 1 and 2 with version 3 of the xGen™ Hybridization and Wash Kit?
No. Version 3 has different formulations for the buffers than previous versions. Only the xGen Bead Wash buffer is the same as the previous versions. The protocol versions are also not interchangeable.
What is the impact on performance if only using 100 ng of library input with the xGen™ Hybridization and Wash v3 Kit?
Our standard input from versions 1 and 2 can still apply here, 500 ng of library. However, a user can reduce the amount to 100 ng and still expect a high on-target rate. Some other metrics may be variable due to the decreased library complexity.
I switched to xGen™ Hybridization and Wash v3 Kit and noticed a shift in my GC read distribution. Is this normal?
xGen Hybridization Wash v3 Kit will produce an improved AT/GC balance to better reflect the probe panel design and library input content. The buffer compositions have changed to minimize bias from hybridization, capture, and wash. Therefore, most users who switch from v1 & v2 to v3 may observe an increase in AT content relative to the GC content.
The post-capture AT vs. GC content is largely dictated by the input library and the panel. Therefore, the resulting GC content would be a combination of the interactions between the library, the panel, the workflow, and the post-capture amplification mix.
Does xGen™ Hybridization and Wash v3 Kit performance benefit from longer hybridization incubation?
Longer hybridization incubation would result in less variability, higher on-target, and higher post-capture library diversity. Increasing from 1-hour to 2-hour hybridization can yield significant positive differences. The impact on results will stem from the library, capture panel, and their interactions.
Our lab uses non-IDT library preps that worked well with xGen™ Hybridization and Wash v1 and v2 Kits. Can we use the xGen Hybridization and Wash v3 Kit for these libraries?
The new kit is compatible with libraries that worked with xGen Hybridization and Wash v1 and v2 Kits.
What types of libraries can be used with xGen™ Hybridization and Wash v3 Kit?
Users can expect libraries that were used with previous versions to be enriched with version 3 of the kit. We recommend libraries generated with xGen library preparation kits such as:
- xGen cfDNA and FFPE DNA Library Prep Kit
- xGen DNA Library Prep Kit EZ & EZ UNI
- xGen DNA Library prep MC & MC UNI
- xGen RNA and Broad Range Kit
What are the buffer storage conditions for the xGen™ Hybridization and Wash v3 Kit?
Reagents are shipped on dry ice except the xGen Hybridization and Wash Beads. Wash Buffers (Bead, Stringent, Buffer I, Buffer II) and Capture Beads can be stored at 4°C. Please store the xGen Hybridization buffer, Cot-1 DNA, and the xGen 2X HiFI PCR master mix at −20°C. Allow all buffers to come to room temperature before using.
What are some of the considerations with a short hybridization time?
If a user would like to use a short hybridization time (1 to 2 hours), optimization may be needed. Results may be variable and will depend on the capture panel (design and target size), desired analysis, and sample type.
The recommended library input range is 100 ng to 6 µg of total library. Target input will be enriched, however, we recommend not exceeding 2.5 µg total library input for 1-hour hybridization times.
What is new about version 3 of the xGen™ Hybridization and Wash Kit?
Version 3 can be used in the same manner as previous versions of the xGen Hybridization and Wash kits. IDT has optimized the buffers to reduce the overall number of washing steps and ease the workflow. Customers are not required to change their xGen capture panels or libraries than previously used.
We also introduce an optional short hybridization period of only 1 hour. Customers can still use the standard 4- or 16-hour incubations. We recommend users empirically test the shorter hybridization time for their workflows.
We have also lowered the supported input for hybridization capture to 100 ng of library instead of 500 ng.
References
- Blumenstiel B, Cibulskis K, et al. Targeted exon sequencing by in-solution hybrid selection. Curr Protoc Hum Genet. 2010;Chapter 18:Unit 18.4.
- Hodges E, Rooks M, et al. Hybrid selection of discrete genomic intervals on custom-designed microarrays for massively parallel sequencing. Nat Protoc. 2009;4(6):960-974.
TruSeq® and Nextera® are registered trademarks of Illumina, Inc., and used with permission. All rights reserved.